Using Social Networking to Learn Genetics
The rise of social networking is perhaps the most spectacular—and to many the most unexpected—phenomenon of the 21st century. It continues to have dramatic political and cultural effects around the globe and its impact on commerce is unprecedented. Seventy-seven percent of youth aged 12 to 17 in the United States are a member of at least one social network1.
All this raises questions for educators. How can social networking best be adapted to create effective learning environments? How broad an impact is this new kind of learning environment likely to have? Can students learn important concepts and skills simply by interacting in a shared virtual environment? How can we assess what they have learned? What are the barriers to using this novel learning modality in schools, and how can they be overcome?
The GeniVille project, funded by the National Science Foundation, was aimed at obtaining tentative answers to questions like these. We integrated our popular genetics software—Geniverse—into a social networking environment called Whyville, which is intended to give tweens aged 8 to 15 opportunities to learn STEM concepts through collaboration and exploration. The result of this union is called Dragons in Whyville.
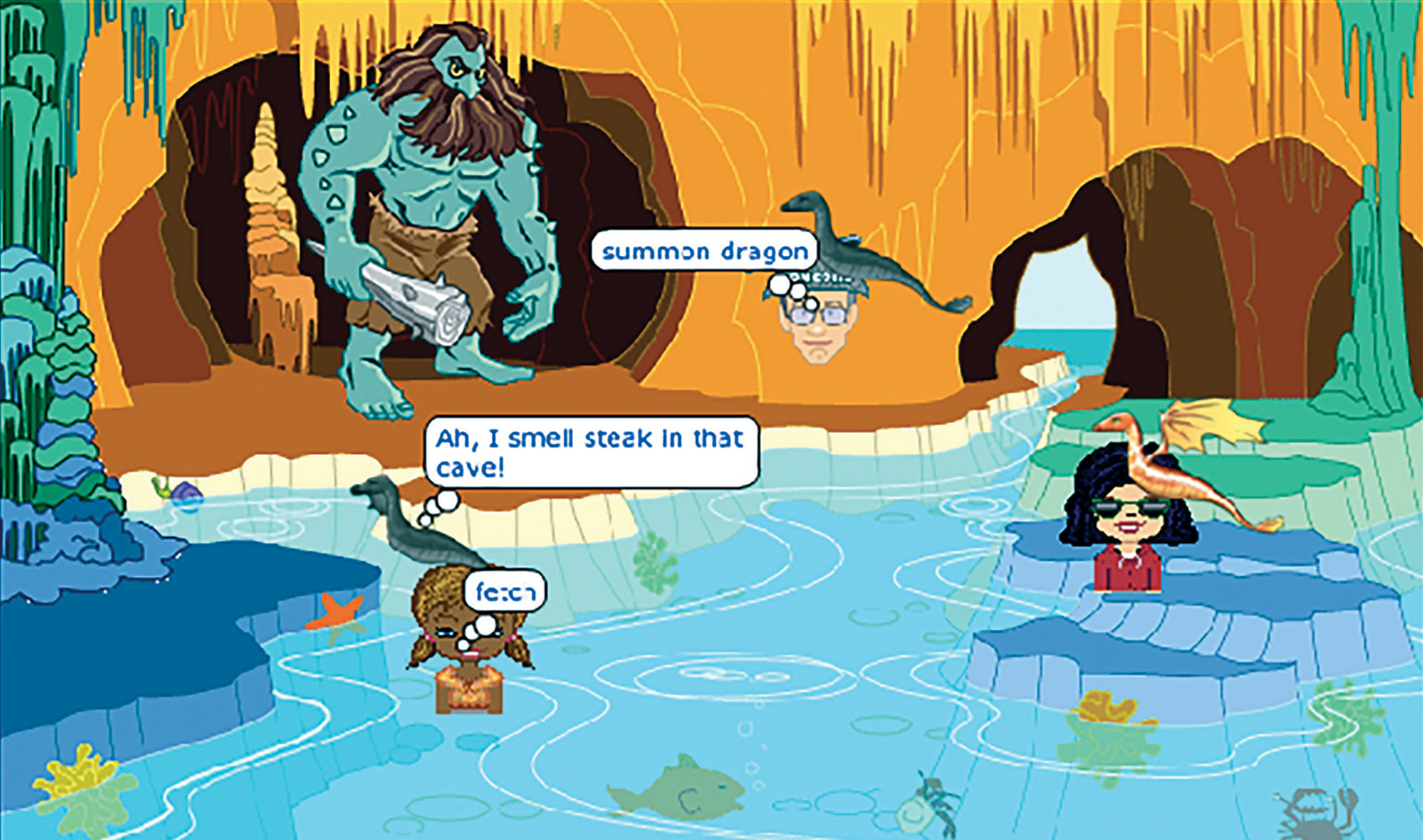

The Dragons game
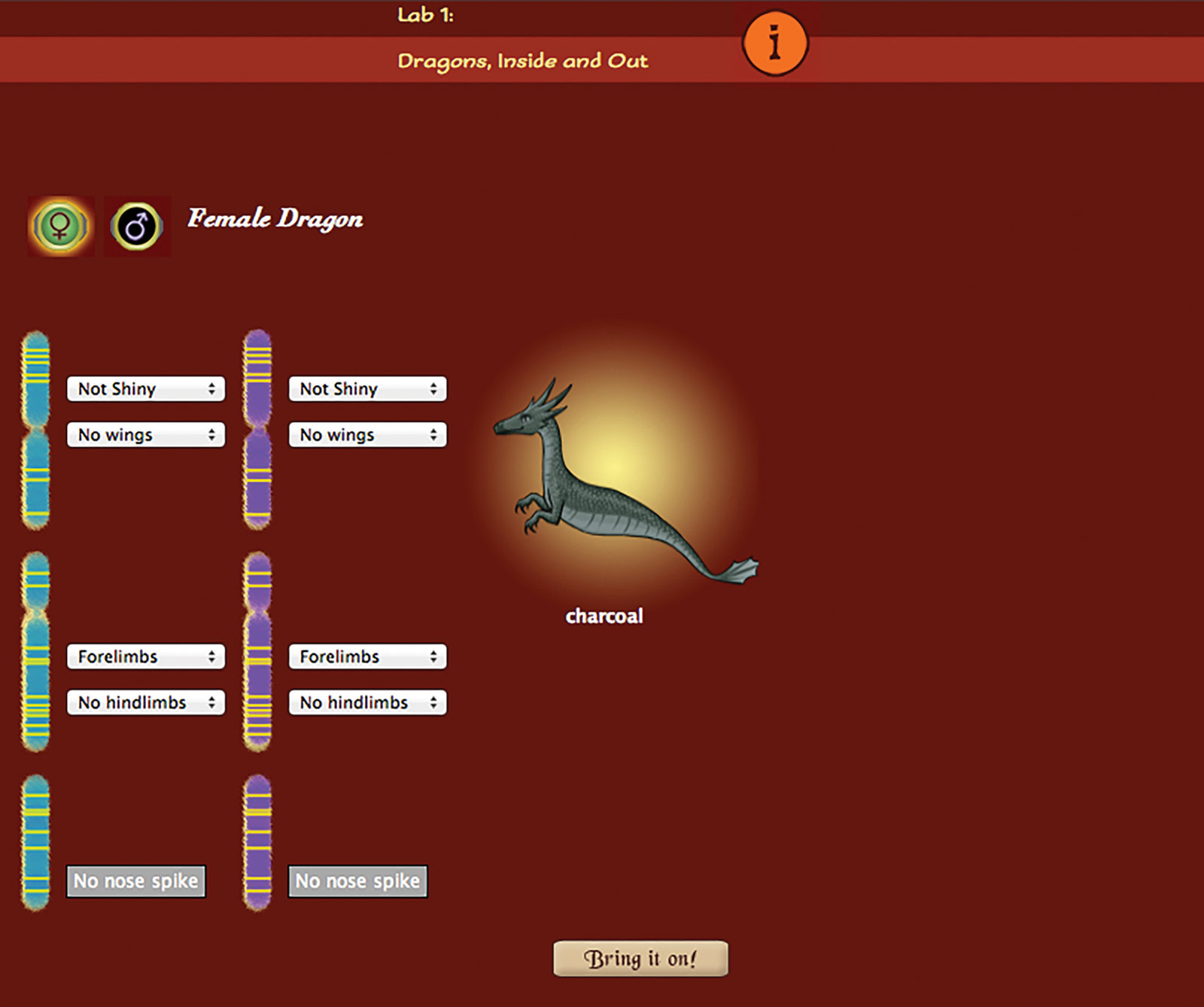
To play the game, users enter the Dragon Castle in Whyville. They visit lairs (where wild dragons roam) to solve breeding challenges or go to labs to delve into dragon traits and inheritance. Sixteen challenges present progressively more complex patterns of inheritance, such as incomplete dominance, sex-linkage and polyallelic traits. Each lair challenge focuses on a specific concept and has one or more labs associated with it. Twenty-one labs provide highly scaffolded activities that highlight genotypic to phenotypic relationships, the processes of meiosis and fertilization and the selection of parents based on the need for certain alleles in a pool of offspring.
For the lair challenges, players must produce a dragon with specific traits that will enable it to fetch treasure. For instance, to find a dragon that can fly to the top of a palm tree and grab the golden coconut, players must enter a lair where dragons have the alleles needed to produce wings and arms. Using a special tool, they can “scope” a dragon or an egg to reveal its chromosomes and alleles. This is helpful in deciding, for instance, whether or not two dragons should be bred or if a particular egg should be hatched. While the labs are single-player activities, players can work alone or collaborate in the lairs to breed offspring for the required traits.
Dragons in Whyville and in school
We released the game to the general Whyville population in September 2013 with 10 challenges and 13 labs, plus a pre-test and post-test. In November 2013, we added a second set of six challenges and eight labs that involved more complex forms of inheritance. Finally, in April 2014, we introduced a fast-paced, multi-player or single-player “egg game,” designed to encourage students to think about the genotypes of eggs before hatching them.
Since we also wanted to gauge the utility of Dragons in a school environment, in June 2014 we invited four teachers at a local, suburban middle school to participate. After completing their standard genetics unit, 455 eighth grade biology students used Dragons during class time for three consecutive days.
What did we learn?
In the year following the introduction of Dragons, over 140,000 individuals logged into Whyville on a minimum of 10 separate occasions. Of these, 8,460 interacted with the game, about half completing only the pre-test (presumably to receive the Whyville currency of “clams” for doing so). Only 356 players took both the pre- and post-tests.
Of the 455 students who participated in Dragons as part of the school implementation, 311 took both the pre-test and the post-test. (Some students did not complete the minimum of six activities required to unlock the post-test, which accounts for the discrepancy.)
The pre- and post-tests were identical and consisted of 11 questions. Five were designed to characterize the players’ attitudes toward genetics and to provide a measure of “self-efficacy” by eliciting their opinions regarding their knowledge of genetics. Students were asked to rate statements on a five-point Likert scale with responses ranging from “Strongly agree,” which we scored as 5 to “Strongly disagree,” scored as 1. In both the “Whyville at large” population and the school population we found no significant differences between the pre- and the post-test on these questions except for two of them.
The only statements on which players changed their answers between the pre-test and the post-test to a statistically significant degree are those that measure self-efficacy, and in both cases students’ confidence in their knowledge of genetics increased. Although the differences in the scores are small, they are highly significant for the pooled (Whyville plus school) data, as shown in Table 12.
| How true is this statement? | pre-test mean score | post-test mean score | p-value of difference (two-tailed paired t-test, n = 667) |
|---|---|---|---|
| 1. I’m interested in learning about genetics. | 3.63 | 3.63 | .50 |
| 2. I like genetics. | 3.53 | 3.59 | .056 |
| 3. I think genetics is useful. | 4.17 | 4.17 | .79 |
| 4. I am good at genetics. | 3.50 | 3.64 | .0026 |
| 5. I know more about genetics than my friends. | 3.08 | 3.19 | .0006 |
Table 1. Comparison of answers to questions on pre- and post-test (all data combined).
In addition to the pre- and post-test data, we also collected a vast amount of performance data, consisting of time-stamped logs of every action taken by every dragon game player. We can determine, for instance, how many lairs each player entered, how many labs they completed and how many challenges they succeeded at. We can even compute how often they used the scoping tool. Since scoping shows the chromosomes and alleles of dragons and eggs, use of this tool is an indication that the player is thinking genotypically and not concentrating solely on the macroscopic appearance of organisms (phenotypic thinking).
Analysis of the school data demonstrates that students shifted from scoping dragons to scoping eggs over the course of their three-day interaction with Dragons. We interpret this unanticipated finding as an indication that students are becoming increasingly conscious of the importance of genes as determinants of a dragon’s traits—a sign that they are learning to think like geneticists. We have not yet analyzed the data from the general Whyville population to see whether they followed a similar learning path.
Our experience in the classroom uncovered a number of barriers that must be faced before the game can be used effectively in that environment. The great value of Whyville—the fact that it is such a rich and engaging environment—means that it offers many distractions to students, making it difficult to keep them focused on the task at hand, thus exacerbating a classroom management problem familiar to every teacher. Technological fixes will have to be in place before the bridge can be crossed between social networking and formal education.
Though our data analysis is not yet complete, we are pleased to note that many kids seem to have enjoyed Dragons, and that their interaction with it has had a positive effect on their self-reported confidence in their knowledge of genetics. It may also have increased their ability to reason like geneticists, though confirmation of this will have to await further analysis.
1 http://www.pewinternet.org/fact-sheets/teens-fact-sheet/
2 Both questions show significant gains for the school data alone, but only Question 5 shows significant gains for the Whyville data alone.
Paul Horwitz (phorwitz@concord.org) directs the GeniVille project.
This material is based upon work supported by the National Science Foundation under grant DRL-1238625. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of the National Science Foundation.